Despite enormous progress over the past century, cancer is still something that ends life. Why then should cancer researchers care about the polar opposite? Why should we care about the origins of life?
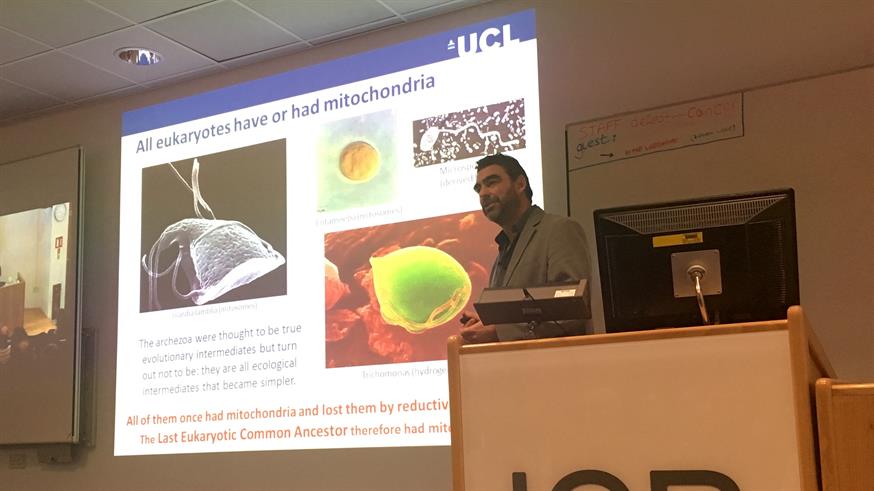
This was the challenge facing UCL’s Dr Nick Lane as he presented the ICR’s annual ‘Darwin Lecture’ last Thursday — 208 years (to the week) from Charles Darwin’s birth. With an enigmatic title of ‘The Origins of Life on Planet Earth’, Dr Lane had attracted teeming audiences in both Chelsea and Sutton. Both lecture theatres were standing room only. Cancer biologists, it seems, are rather interested in such big questions.
Dr Lane started by describing the great ‘evolutionary scandal’ at the heart of biology. Why, he asked, do we not see eukaryotic traits in prokaryotes? Why did eukaryotic complexity not co-evolve in bacteria? The answer, he revealed, lies in bioenergetics. Dr Lane explained how eukaryotic life originated from a single ‘big bang’ event 1.5–2.0 billion years ago, when an archaeon engulfed a proteobacteria. This endosymbiosed aerobic bacteria enabled the host archaea to massively increase its energy production — providing the metabolic resources to develop further complexity. Eukaryotes had arisen not from a linear evolution of one cell type, but from a combination of different cells. Less of a ‘tree of life’ and more a ‘ring of life’. Empowered by their newfound energy, early eukaryotes eventually evolved into the pantheon of multicellular biology we see today. It was a humbling perspective of our cellular origins.
I think it’s fair to say that ICR scientists rarely consider such topics. In many ways, ‘origin of life’ research is the ultimate blue sky endeavour — studied because it’s fascinating, not because it’s obviously useful. This research is somewhat removed from the ‘translational research’ the ICR aspires to. Yet the abundant audiences across both ICR sites demonstrated there is a real craving for such insight among translational scientists. There is something universally intriguing about our ancient common ancestors.
Implications for cancer research
So aside from being genuinely fascinating, how does the ‘origin of life’ relate to our day-to-day research of cancer? What can we learn from our cellular history?
From speaking to colleagues after the lecture, everyone took away something different. The genetically inclined were fascinated by how mitochondria lost 99% of their genes just so they could spend the saved ATP on something new. What evolutionary advantages might be afforded by gene loss frequently seen in cancer cells?
The metabolically enthused certainly enjoyed seeing bioenergetics at the centre of cellular evolution. Given metabolic changes are powerful enough to drive the entire evolution of eukaryotic life, maybe we shouldn’t be surprised by its frequent deregulation in cancer. Could cancer just be an intra-organism metabolic shift?
As someone interested in the tumour microenvironment, I was charmed to learn how the eukaryotes, with their great potential for complexity, arose not by serial evolution of a single cell, but by a symbiotic collaboration between two different cell types. In cancer, we often see healthy cells (such as fibroblasts, endothelial, and immune cells) coerced by cancer cells to provide tumours with resources cancer cells can’t make themselves. It’s fascinating to think our eukaryotic ancestors were formed by a similar collaboration of different cells. It implies the improved ‘fitness’ afforded by heterocellular collaboration is both ancient and universal.
Whatever you took away from the lecture, I think it’s fair to say everyone left with a new view on our cellular origins and their consciousness substantially raised. I’m already looking forward to Darwin’s 209th.
This post originally appeared as a 'From Bench To Blog' blog for The Institute of Cancer Research.